Linker Information
General Information of This Linker
| Linker ID |
LIN00117
|
|||||
|---|---|---|---|---|---|---|
| Linker Name |
3,3-Dimethylglutaric acid
|
|||||
| Linker Type |
Enzyme-sensitive linkers
|
|||||
| Structure |
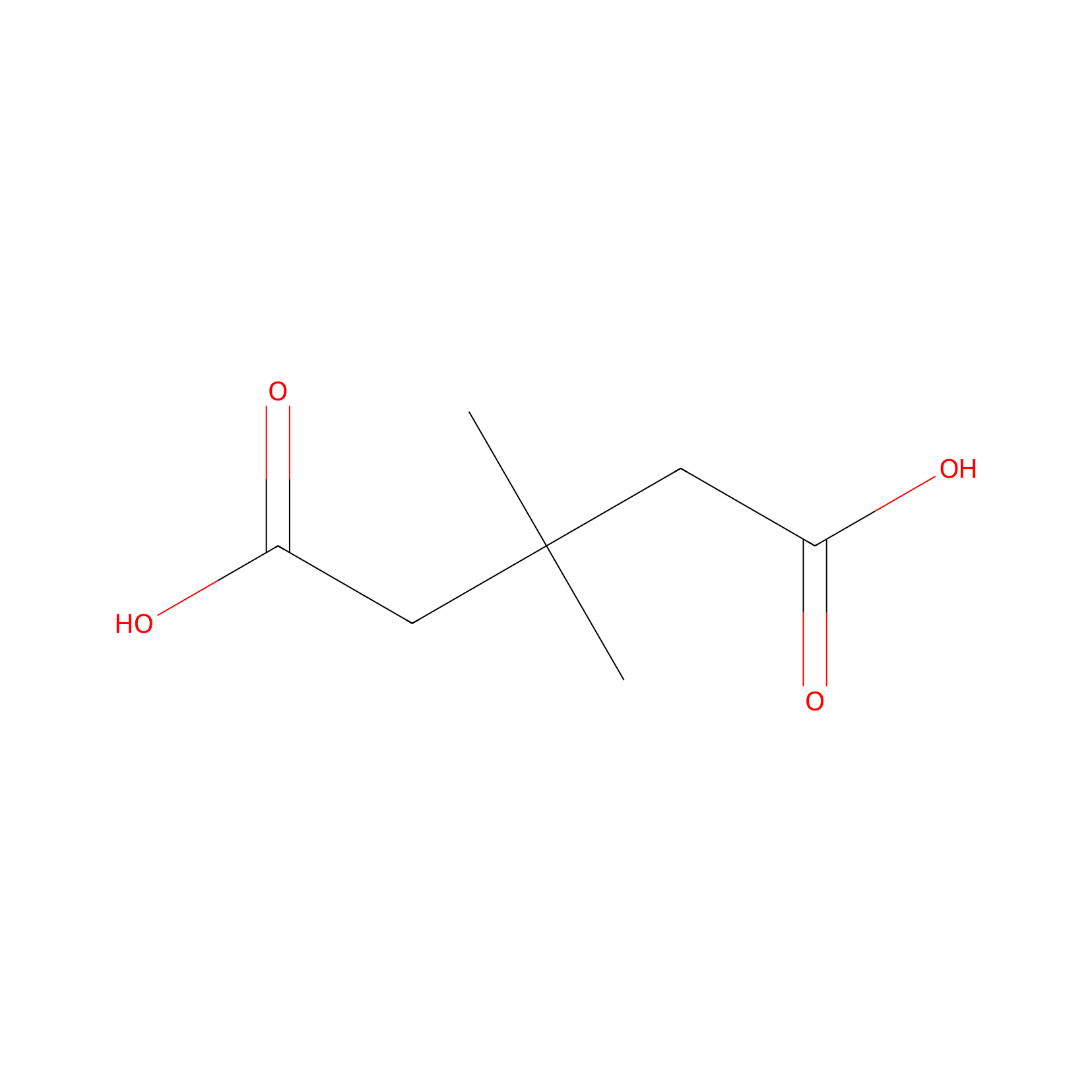
|
|||||
| Formula |
C7H12O4
|
|||||
| #Ro5 Violations (Lipinski): 0 | Molecular Weight (mw) | 160.17 | ||||
| Lipid-water partition coefficient (xlogp) | 0.4 | |||||
| Hydrogen Bond Donor Count (hbonddonor) | 2 | |||||
| Hydrogen Bond Acceptor Count (hbondacc) | 4 | |||||
| Rotatable Bond Count (rotbonds) | 4 | |||||
| PubChem CID | ||||||
| Canonical smiles |
CC(C)(CC(=O)O)CC(=O)O
|
|||||
| InChI |
InChI=1S/C7H12O4/c1-7(2,3-5(8)9)4-6(10)11/h3-4H2,1-2H3,(H,8,9)(H,10,11)
|
|||||
| InChIKey |
DUHQIGLHYXLKAE-UHFFFAOYSA-N
|
|||||
| IUPAC Name |
3,3-dimethylpentanedioic acid
|
|||||
Each Peptide-drug Conjugate Related to This Linker
Full Information of The Activity Data of The PDC(s) Related to This Linker
TH1904 [Preclinical]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Ovarian cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
54 nM
|
|||
| Administration Time | 12 h | ||||
| MOA of PDC |
Overall, these results indicate that both conjugates alter cell migration in either TNBC or ovarian cancer cell models through a SORT1-dependent mechanism. The effects of TH1902 and TH1904 on cell migration further support the molecular rationale that inhibition of VM activity may result, in part, from the reduction of cancer cell invasive phenotype.
Click to Show/Hide
|
||||
| Description |
TH1904 IC50 value was 54 nM for the number of loops inhibition (Figure 5B).
|
||||
| In Vitro Model | Ovarian clear cell adenocarcinoma | ES-2 cell | CVCL_3509 | ||
References