Peptide Information
General Information of This Peptide
| Peptide ID |
PEP00017
|
|||||
|---|---|---|---|---|---|---|
| Peptide Name |
Peptide 1131
|
|||||
| Structure |
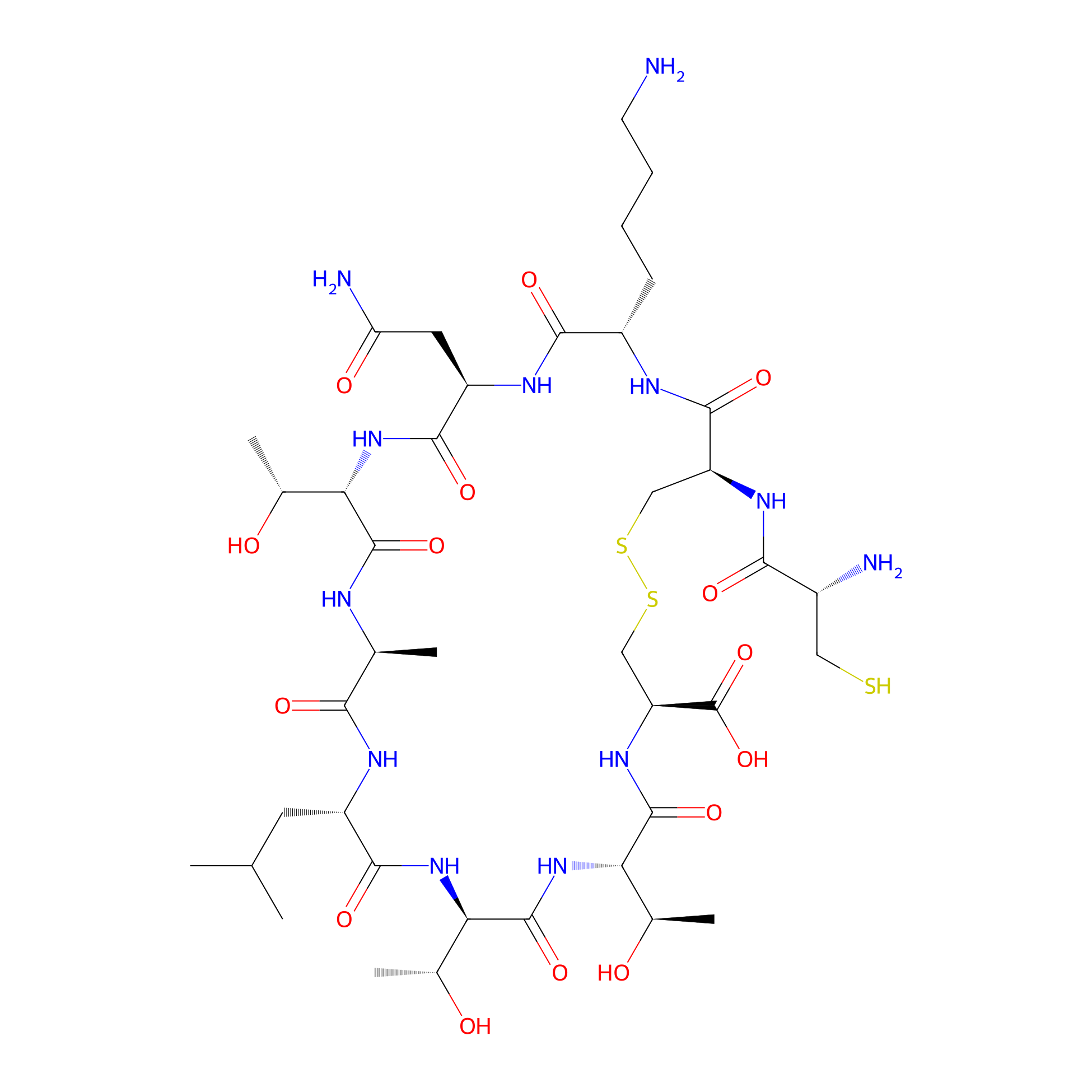
|
|||||
| Sequence |
CKNTALTTC
|
|||||
| Peptide Type |
Cyclic
|
|||||
| Receptor Name |
Kita-kyushu lung cancer antigen 1 (CT83)
|
Receptor Info | ||||
| PDC Transmembrane Types | Cell-penetrating peptides (CPPs) | |||||
| Formula |
C40H70N12O15S3
|
|||||
| Isosmiles |
[H]N[C@H](CS)C(=O)N[C@H]1CSSC[C@@H](C(=O)O)NC(=O)[C@]([H])([C@@H](C)O)NC(=O)[C@@]([H])([C@@H](C)O)NC(=O)[C@H](CC(C)C)NC(=O)[C@H](C)NC(=O)[C@]([H])([C@@H](C)O)NC(=O)[C@@H](CC(N)=O)NC(=O)[C@H](CCCCN)NC1=O
|
|||||
| InChI |
InChI=1S/C40H70N12O15S3/c1-16(2)11-23-34(60)51-30(20(6)55)39(65)52-29(19(5)54)38(64)49-26(40(66)67)15-70-69-14-25(48-32(58)21(42)13-68)36(62)45-22(9-7-8-10-41)33(59)47-24(12-27(43)56)35(61)50-28(18(4)53)37(63)44-17(3)31(57)46-23/h16-26,28-30,53-55,68H,7-15,41-42H2,1-6H3,(H2,43,56)(H,44,63)(H,45,62)(H,46,57)(H,47,59)(H,48,58)(H,49,64)(H,50,61)(H,51,60)(H,52,65)(H,66,67)/t17-,18+,19+,20+,21+,22-,23-,24+,25-,26-,28-,29-,30+/m0/s1
|
|||||
| InChIKey |
FFRNBDGZFKDTON-OKBHEYPESA-N
|
|||||
| Pharmaceutical Properties |
Molecule Weight
|
1055.27
|
Polar area
|
455.02
|
||
|
Complexity
|
1054.424573
|
xlogp Value
|
-6.7009
|
|||
|
Heavy Count
|
70
|
Rot Bonds
|
16
|
|||
|
Hbond acc
|
19
|
Hbond Donor
|
17
|
|||
The Activity Data of This Peptide
| Peptide Activity Information 1 | [1] | |||||
| Fluorescence intensity | 2000 | |||||
|---|---|---|---|---|---|---|
| Binding Affinity Assay |
For the fluorescence-linked immunosorbent assay (FLISA), flat-bottom 96-well cell culture plates (Thermo Fisher Scientific, Waltham, MA, USA) were coated with 10 μg/mL KK-LC-1 protein solution at 4 °C overnight and then blocked with 1% BSA-PBS at room temperature for 1 h. The plates were washed with PBS containing 0.1% Tween 20 (0.1% PBST). Three-fold dilutions of FITC-labeled 1131-MMAE, 1131 peptide and the non-targeting control CG7C-MMAE were added and incubated at room temperature for 2 h. The plates were washed again and the fluorescence intensity was measured with a multimode microplate reader (Tecan, Mendov, Switzerland). The excitation wavelength was 485 nm and emission wavelength was 535 nm.
Click to Show/Hide
|
|||||
| Experimental Condition | 10 μg/mL KK-LC-1 protein solution at 4 C overnight | |||||
Each Peptide-drug Conjugate Related to This Peptide
Full Information of The Activity Data of The PDC(s) Related to This Peptide
1131-MMAE [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Gastric cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
3.87 nM
|
|||
| Administration Time | 72 h | ||||
| Description |
We next evaluated the in vitro cytotoxicity of 1131-MMAE. NUGC-4, MKN45 and HGC27 cells were drug-treated for 72 h and their viability was assessed. 1131-MMAE killed KK-LC-1 positive gastric cancer cells with high potency. The IC50 values of 1131-MMAE were 3.87 nM for NUGC-4 cells and 5.26 nM for MKN45 cells. However, although a very high concentration was used, 1131-MMAE could only moderately inhibit the viability of KK-LC-1 negative HGC27 cells. The IC50 value of 1131-MMAE for HGC27 cells was 100-200 times higher than that in NUGC-4 and MNK45 cells. Free MMAE exerted cytotoxicity irrespective of the KK-LC-1 expression status. The IC50 values of free MMAE were 10.97 nM for NUGC-4 cells, 10.70 nM for MKN45 cells and 7.18 nM for HGC27 cells. Naked 1131 peptide showed no cytotoxicity to all three cell lines. These results were consistent with the KK-LC-1-dependent endocytosis and confirmed the target-selective killing of 1131-MMAE.
Click to Show/Hide
|
||||
| In Vitro Model | Gastric signet ring cell adenocarcinoma | NUGC-4 cell | CVCL_3082 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Gastric cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
5.26 nM
|
|||
| Administration Time | 72 h | ||||
| Description |
We next evaluated the in vitro cytotoxicity of 1131-MMAE. NUGC-4, MKN45 and HGC27 cells were drug-treated for 72 h and their viability was assessed. 1131-MMAE killed KK-LC-1 positive gastric cancer cells with high potency. The IC50 values of 1131-MMAE were 3.87 nM for NUGC-4 cells and 5.26 nM for MKN45 cells. However, although a very high concentration was used, 1131-MMAE could only moderately inhibit the viability of KK-LC-1 negative HGC27 cells. The IC50 value of 1131-MMAE for HGC27 cells was 100-200 times higher than that in NUGC-4 and MNK45 cells. Free MMAE exerted cytotoxicity irrespective of the KK-LC-1 expression status. The IC50 values of free MMAE were 10.97 nM for NUGC-4 cells, 10.70 nM for MKN45 cells and 7.18 nM for HGC27 cells. Naked 1131 peptide showed no cytotoxicity to all three cell lines. These results were consistent with the KK-LC-1-dependent endocytosis and confirmed the target-selective killing of 1131-MMAE.
Click to Show/Hide
|
||||
| In Vitro Model | Gastric adenocarcinoma | MKN45 cell | CVCL_0434 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Gastric cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
786 nM
|
|||
| Administration Time | 72 h | ||||
| Description |
We next evaluated the in vitro cytotoxicity of 1131-MMAE. NUGC-4, MKN45 and HGC27 cells were drug-treated for 72 h and their viability was assessed. 1131-MMAE killed KK-LC-1 positive gastric cancer cells with high potency. The IC50 values of 1131-MMAE were 3.87 nM for NUGC-4 cells and 5.26 nM for MKN45 cells. However, although a very high concentration was used, 1131-MMAE could only moderately inhibit the viability of KK-LC-1 negative HGC27 cells. The IC50 value of 1131-MMAE for HGC27 cells was 100-200 times higher than that in NUGC-4 and MNK45 cells. Free MMAE exerted cytotoxicity irrespective of the KK-LC-1 expression status. The IC50 values of free MMAE were 10.97 nM for NUGC-4 cells, 10.70 nM for MKN45 cells and 7.18 nM for HGC27 cells. Naked 1131 peptide showed no cytotoxicity to all three cell lines. These results were consistent with the KK-LC-1-dependent endocytosis and confirmed the target-selective killing of 1131-MMAE.
Click to Show/Hide
|
||||
| In Vitro Model | Gastric carcinoma | HGC-27 cell | CVCL_1279 | ||
References