Peptide Information
General Information of This Peptide
| Peptide ID |
PEP01099
|
|||||
|---|---|---|---|---|---|---|
| Peptide Name |
WX(D)EAAYQKFL peptide
|
|||||
| Structure |
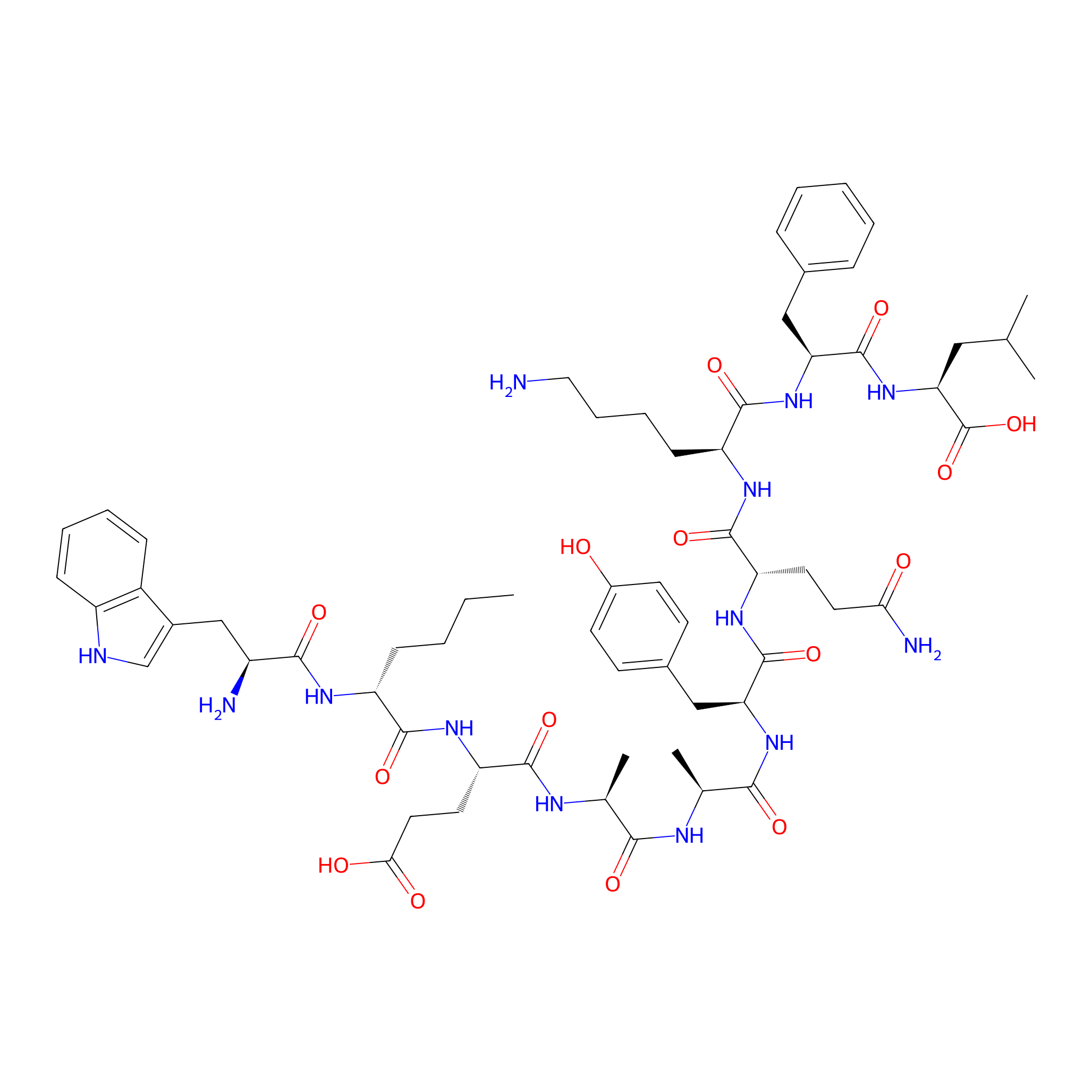
|
|||||
| Sequence |
WX(D)EAAYQKFL
|
|||||
| Peptide Type |
Linear
|
|||||
| Receptor Name |
Keratin, type II cytoskeletal 1 (KRT1)
|
Receptor Info | ||||
| PDC Transmembrane Types | Cell targeting peptides (CTPs) | |||||
| Formula |
C63H89N13O15
|
|||||
| Isosmiles |
[H]NC(=O)CC[C@H](NC(=O)[C@H](Cc1ccc(O[H])cc1)NC(=O)[C@H](C)NC(=O)[C@H](C)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@@H](CCCC)NC(=O)[C@@H](N)Cc1cn([H])c2ccccc12)C(=O)N[C@@H](CCCCN)C(=O)N[C@@H](Cc1ccccc1)C(=O)N[C@@H](CC(C)C)C(=O)O
|
|||||
| InChI |
InChI=1S/C63H89N13O15/c1-6-7-18-45(70-56(83)43(65)33-40-34-67-44-19-12-11-17-42(40)44)58(85)72-48(26-28-53(79)80)57(84)69-36(4)54(81)68-37(5)55(82)74-49(32-39-21-23-41(77)24-22-39)61(88)73-47(25-27-52(66)78)60(87)71-46(20-13-14-29-64)59(86)75-50(31-38-15-9-8-10-16-38)62(89)76-51(63(90)91)30-35(2)3/h8-12,15-17,19,21-24,34-37,43,45-51,67,77H,6-7,13-14,18,20,25-33,64-65H2,1-5H3,(H2,66,78)(H,68,81)(H,69,84)(H,70,83)(H,71,87)(H,72,85)(H,73,88)(H,74,82)(H,75,86)(H,76,89)(H,79,80)(H,90,91)/t36-,37-,43-,45+,46-,47-,48-,49-,50-,51-/m0/s1
|
|||||
| InChIKey |
RCEMGKLFRIOKFG-GYMLMKCOSA-N
|
|||||
| Pharmaceutical Properties |
Molecule Weight
|
1268.481
|
Polar area
|
467.65
|
||
|
Complexity
|
1267.660109
|
xlogp Value
|
0.2119
|
|||
|
Heavy Count
|
91
|
Rot Bonds
|
41
|
|||
|
Hbond acc
|
15
|
Hbond Donor
|
16
|
|||
Each Peptide-drug Conjugate Related to This Peptide
Full Information of The Activity Data of The PDC(s) Related to This Peptide
FITC-WX(D)EAAYQKFL peptide conjugate [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Mean fluorescence intensities |
25
|
|||
| Administration Time | 30 min | ||||
| Administration Dosage | 1 μM | ||||
| Evaluation Method | LSR Fortessa flow cytometer | ||||
| MOA of PDC |
Here we have designed cyclic analogues of peptide4to enhance the affinity and specificity toward breast cancer cell lines while maintaining the proteolytic stability. In addition, we have reduced the number ofd-amino acids from two to one or none, as it is hypothesized that cyclization may impart sufficient stability to the peptide structure. The results show that the cyclic peptide analogues display higher uptake by breast cancer cells than all other analogues tested so far, and show minimal uptake by the noncancerous cells. One of the cyclic analogues with oned-amino acid (7) was sufficiently stable toward proteolytic degradation. When immobilized on gold microcantilever surface, the peptide was able to capture breast cancer cells specifically. Preliminary animal studies using mice carrying orthotopic breast MDA-MB-231 tumors were also performed to track the Cy5.5 labeled peptide7in live mice. The study highlights the discovery of a novel proteolytically stable cyclic peptide7for breast cancer targeting that can be used for targeted drug delivery or for capturing circulating breast tumor cells from human blood samples.
Click to Show/Hide
|
||||
| Description |
Next, we compared the breast cancer cell uptake of the two cyclic peptides,7and8(with alll-amino acids), versus the noncancerous cell uptake to three previously reported lead sequences (linear peptides), namely,1,2, and4. Two peptide concentrations were used, 1 μM and 10 μM. As shown inFigure S3, the magnitude of cell uptake increased considerably at higher peptide concentration (10 μM). The intensity of cell-associated fluorescence at 10 μM peptide concentration was at least 2-fold higher when compared to the cell-associated fluorescence observed at 1 μM peptide concentration. This trend was observed for all breast cancer cell lines used in the study. The mean fluorescence intensities (MFIs) of FITC-7at 10 μM were 2771 ± 78, 2607 ± 116, and 1761 ± 83 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively, whereas the MFIs were 179 ± 25 and 247 ± 27, respectively, when incubated with normal HUVEC and MCF10A. In comparison, the MFIs for the linear peptide with twod-amino acids x and k (FITC-4) were 965 ± 44, 1666 ± 97, and 1031 ± 54 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively. Overall the uptake was observed in the following order1<2<4<7<8with peptide8displaying highest cellular uptake. Noncancerous cell uptake (HUVEC and breast tissue derived MCF-10A cells) of all the peptides was minimal. Even at high peptide concentration (10 μM) the uptake of the peptides was low suggesting the presence of smaller number of putative receptors on these cells.
Click to Show/Hide
|
||||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Mean fluorescence intensities |
25
|
|||
| Administration Time | 30 min | ||||
| Administration Dosage | 1 μM | ||||
| Evaluation Method | LSR Fortessa flow cytometer | ||||
| MOA of PDC |
Here we have designed cyclic analogues of peptide4to enhance the affinity and specificity toward breast cancer cell lines while maintaining the proteolytic stability. In addition, we have reduced the number ofd-amino acids from two to one or none, as it is hypothesized that cyclization may impart sufficient stability to the peptide structure. The results show that the cyclic peptide analogues display higher uptake by breast cancer cells than all other analogues tested so far, and show minimal uptake by the noncancerous cells. One of the cyclic analogues with oned-amino acid (7) was sufficiently stable toward proteolytic degradation. When immobilized on gold microcantilever surface, the peptide was able to capture breast cancer cells specifically. Preliminary animal studies using mice carrying orthotopic breast MDA-MB-231 tumors were also performed to track the Cy5.5 labeled peptide7in live mice. The study highlights the discovery of a novel proteolytically stable cyclic peptide7for breast cancer targeting that can be used for targeted drug delivery or for capturing circulating breast tumor cells from human blood samples.
Click to Show/Hide
|
||||
| Description |
Next, we compared the breast cancer cell uptake of the two cyclic peptides,7and8(with alll-amino acids), versus the noncancerous cell uptake to three previously reported lead sequences (linear peptides), namely,1,2, and4. Two peptide concentrations were used, 1 μM and 10 μM. As shown inFigure S3, the magnitude of cell uptake increased considerably at higher peptide concentration (10 μM). The intensity of cell-associated fluorescence at 10 μM peptide concentration was at least 2-fold higher when compared to the cell-associated fluorescence observed at 1 μM peptide concentration. This trend was observed for all breast cancer cell lines used in the study. The mean fluorescence intensities (MFIs) of FITC-7at 10 μM were 2771 ± 78, 2607 ± 116, and 1761 ± 83 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively, whereas the MFIs were 179 ± 25 and 247 ± 27, respectively, when incubated with normal HUVEC and MCF10A. In comparison, the MFIs for the linear peptide with twod-amino acids x and k (FITC-4) were 965 ± 44, 1666 ± 97, and 1031 ± 54 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively. Overall the uptake was observed in the following order1<2<4<7<8with peptide8displaying highest cellular uptake. Noncancerous cell uptake (HUVEC and breast tissue derived MCF-10A cells) of all the peptides was minimal. Even at high peptide concentration (10 μM) the uptake of the peptides was low suggesting the presence of smaller number of putative receptors on these cells.
Click to Show/Hide
|
||||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Mean fluorescence intensities |
80
|
|||
| Administration Time | 30 min | ||||
| Administration Dosage | 1 μM | ||||
| Evaluation Method | LSR Fortessa flow cytometer | ||||
| MOA of PDC |
Here we have designed cyclic analogues of peptide4to enhance the affinity and specificity toward breast cancer cell lines while maintaining the proteolytic stability. In addition, we have reduced the number ofd-amino acids from two to one or none, as it is hypothesized that cyclization may impart sufficient stability to the peptide structure. The results show that the cyclic peptide analogues display higher uptake by breast cancer cells than all other analogues tested so far, and show minimal uptake by the noncancerous cells. One of the cyclic analogues with oned-amino acid (7) was sufficiently stable toward proteolytic degradation. When immobilized on gold microcantilever surface, the peptide was able to capture breast cancer cells specifically. Preliminary animal studies using mice carrying orthotopic breast MDA-MB-231 tumors were also performed to track the Cy5.5 labeled peptide7in live mice. The study highlights the discovery of a novel proteolytically stable cyclic peptide7for breast cancer targeting that can be used for targeted drug delivery or for capturing circulating breast tumor cells from human blood samples.
Click to Show/Hide
|
||||
| Description |
Next, we compared the breast cancer cell uptake of the two cyclic peptides,7and8(with alll-amino acids), versus the noncancerous cell uptake to three previously reported lead sequences (linear peptides), namely,1,2, and4. Two peptide concentrations were used, 1 μM and 10 μM. As shown inFigure S3, the magnitude of cell uptake increased considerably at higher peptide concentration (10 μM). The intensity of cell-associated fluorescence at 10 μM peptide concentration was at least 2-fold higher when compared to the cell-associated fluorescence observed at 1 μM peptide concentration. This trend was observed for all breast cancer cell lines used in the study. The mean fluorescence intensities (MFIs) of FITC-7at 10 μM were 2771 ± 78, 2607 ± 116, and 1761 ± 83 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively, whereas the MFIs were 179 ± 25 and 247 ± 27, respectively, when incubated with normal HUVEC and MCF10A. In comparison, the MFIs for the linear peptide with twod-amino acids x and k (FITC-4) were 965 ± 44, 1666 ± 97, and 1031 ± 54 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively. Overall the uptake was observed in the following order1<2<4<7<8with peptide8displaying highest cellular uptake. Noncancerous cell uptake (HUVEC and breast tissue derived MCF-10A cells) of all the peptides was minimal. Even at high peptide concentration (10 μM) the uptake of the peptides was low suggesting the presence of smaller number of putative receptors on these cells.
Click to Show/Hide
|
||||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Mean fluorescence intensities |
85
|
|||
| Administration Time | 30 min | ||||
| Administration Dosage | 1 μM | ||||
| Evaluation Method | LSR Fortessa flow cytometer | ||||
| MOA of PDC |
Here we have designed cyclic analogues of peptide4to enhance the affinity and specificity toward breast cancer cell lines while maintaining the proteolytic stability. In addition, we have reduced the number ofd-amino acids from two to one or none, as it is hypothesized that cyclization may impart sufficient stability to the peptide structure. The results show that the cyclic peptide analogues display higher uptake by breast cancer cells than all other analogues tested so far, and show minimal uptake by the noncancerous cells. One of the cyclic analogues with oned-amino acid (7) was sufficiently stable toward proteolytic degradation. When immobilized on gold microcantilever surface, the peptide was able to capture breast cancer cells specifically. Preliminary animal studies using mice carrying orthotopic breast MDA-MB-231 tumors were also performed to track the Cy5.5 labeled peptide7in live mice. The study highlights the discovery of a novel proteolytically stable cyclic peptide7for breast cancer targeting that can be used for targeted drug delivery or for capturing circulating breast tumor cells from human blood samples.
Click to Show/Hide
|
||||
| Description |
Next, we compared the breast cancer cell uptake of the two cyclic peptides,7and8(with alll-amino acids), versus the noncancerous cell uptake to three previously reported lead sequences (linear peptides), namely,1,2, and4. Two peptide concentrations were used, 1 μM and 10 μM. As shown inFigure S3, the magnitude of cell uptake increased considerably at higher peptide concentration (10 μM). The intensity of cell-associated fluorescence at 10 μM peptide concentration was at least 2-fold higher when compared to the cell-associated fluorescence observed at 1 μM peptide concentration. This trend was observed for all breast cancer cell lines used in the study. The mean fluorescence intensities (MFIs) of FITC-7at 10 μM were 2771 ± 78, 2607 ± 116, and 1761 ± 83 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively, whereas the MFIs were 179 ± 25 and 247 ± 27, respectively, when incubated with normal HUVEC and MCF10A. In comparison, the MFIs for the linear peptide with twod-amino acids x and k (FITC-4) were 965 ± 44, 1666 ± 97, and 1031 ± 54 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively. Overall the uptake was observed in the following order1<2<4<7<8with peptide8displaying highest cellular uptake. Noncancerous cell uptake (HUVEC and breast tissue derived MCF-10A cells) of all the peptides was minimal. Even at high peptide concentration (10 μM) the uptake of the peptides was low suggesting the presence of smaller number of putative receptors on these cells.
Click to Show/Hide
|
||||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Mean fluorescence intensities |
90
|
|||
| Administration Time | 30 min | ||||
| Administration Dosage | 1 μM | ||||
| Evaluation Method | LSR Fortessa flow cytometer | ||||
| MOA of PDC |
Here we have designed cyclic analogues of peptide4to enhance the affinity and specificity toward breast cancer cell lines while maintaining the proteolytic stability. In addition, we have reduced the number ofd-amino acids from two to one or none, as it is hypothesized that cyclization may impart sufficient stability to the peptide structure. The results show that the cyclic peptide analogues display higher uptake by breast cancer cells than all other analogues tested so far, and show minimal uptake by the noncancerous cells. One of the cyclic analogues with oned-amino acid (7) was sufficiently stable toward proteolytic degradation. When immobilized on gold microcantilever surface, the peptide was able to capture breast cancer cells specifically. Preliminary animal studies using mice carrying orthotopic breast MDA-MB-231 tumors were also performed to track the Cy5.5 labeled peptide7in live mice. The study highlights the discovery of a novel proteolytically stable cyclic peptide7for breast cancer targeting that can be used for targeted drug delivery or for capturing circulating breast tumor cells from human blood samples.
Click to Show/Hide
|
||||
| Description |
Next, we compared the breast cancer cell uptake of the two cyclic peptides,7and8(with alll-amino acids), versus the noncancerous cell uptake to three previously reported lead sequences (linear peptides), namely,1,2, and4. Two peptide concentrations were used, 1 μM and 10 μM. As shown inFigure S3, the magnitude of cell uptake increased considerably at higher peptide concentration (10 μM). The intensity of cell-associated fluorescence at 10 μM peptide concentration was at least 2-fold higher when compared to the cell-associated fluorescence observed at 1 μM peptide concentration. This trend was observed for all breast cancer cell lines used in the study. The mean fluorescence intensities (MFIs) of FITC-7at 10 μM were 2771 ± 78, 2607 ± 116, and 1761 ± 83 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively, whereas the MFIs were 179 ± 25 and 247 ± 27, respectively, when incubated with normal HUVEC and MCF10A. In comparison, the MFIs for the linear peptide with twod-amino acids x and k (FITC-4) were 965 ± 44, 1666 ± 97, and 1031 ± 54 when incubated with MCF-7, MDA-MB-231, and MDA-MB-435, respectively. Overall the uptake was observed in the following order1<2<4<7<8with peptide8displaying highest cellular uptake. Noncancerous cell uptake (HUVEC and breast tissue derived MCF-10A cells) of all the peptides was minimal. Even at high peptide concentration (10 μM) the uptake of the peptides was low suggesting the presence of smaller number of putative receptors on these cells.
Click to Show/Hide
|
||||
References