Peptide Information
General Information of This Peptide
| Peptide ID |
PEP00152
|
|||||
|---|---|---|---|---|---|---|
| Peptide Name |
[D-Lys6]-LH-RH
|
|||||
| Structure |
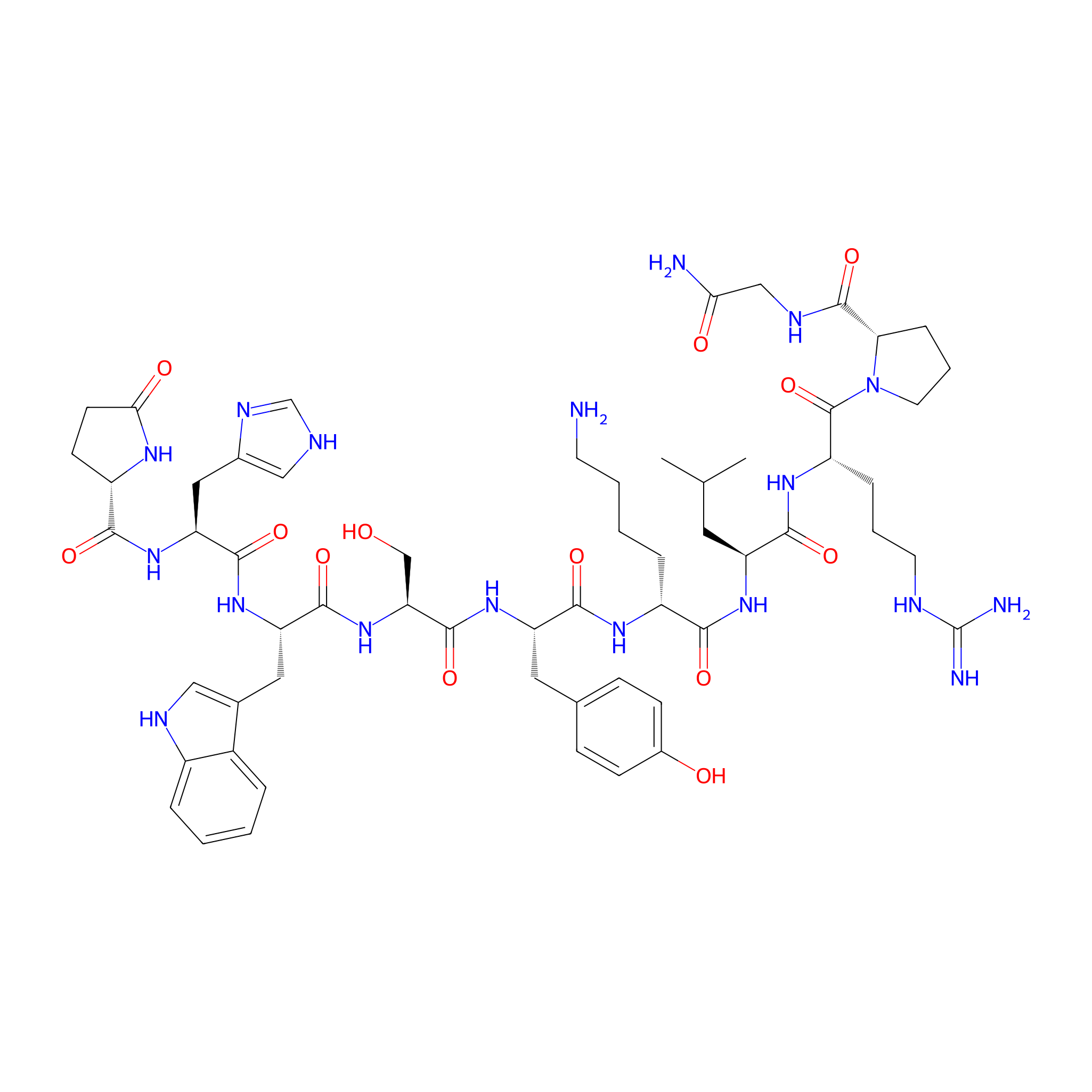
|
|||||
| Sequence |
XHWSYKLRPG
|
|||||
| Peptide Type |
Linear
|
|||||
| Receptor Name |
Gonadotropin-releasing hormone receptor (GNRHR)
|
Receptor Info | ||||
| PDC Transmembrane Types | Cell-penetrating peptides (CPPs) | |||||
| Metabolism |
In most of these studies, [D-Lys6]-LHRH, a degradation-resistant LHRH analog, was used due to its high binding affinity to LHRH-R as well as its reactive site at position 6.
|
|||||
| Formula |
C59H84N18O13
|
|||||
| Isosmiles |
CC(C)C[C@H](NC(=O)[C@@H](CCCCN)NC(=O)[C@H](Cc1ccc(O)cc1)NC(=O)[C@H](CO)NC(=O)[C@H](Cc1c[nH]c2ccccc12)NC(=O)[C@H](Cc1c[nH]cn1)NC(=O)[C@@H]1CCC(=O)N1)C(=O)N[C@@H](CCCNC(=N)N)C(=O)N1CCC[C@H]1C(=O)NCC(N)=O
|
|||||
| InChI |
InChI=1S/C59H84N18O13/c1-32(2)23-42(52(84)71-41(12-7-21-65-59(62)63)58(90)77-22-8-13-47(77)57(89)67-29-48(61)80)72-50(82)39(11-5-6-20-60)70-53(85)43(24-33-14-16-36(79)17-15-33)73-56(88)46(30-78)76-54(86)44(25-34-27-66-38-10-4-3-9-37(34)38)74-55(87)45(26-35-28-64-31-68-35)75-51(83)40-18-19-49(81)69-40/h3-4,9-10,14-17,27-28,31-32,39-47,66,78-79H,5-8,11-13,18-26,29-30,60H2,1-2H3,(H2,61,80)(H,64,68)(H,67,89)(H,69,81)(H,70,85)(H,71,84)(H,72,82)(H,73,88)(H,74,87)(H,75,83)(H,76,86)(H4,62,63,65)/t39-,40+,41+,42+,43+,44+,45+,46+,47+/m1/s1
|
|||||
| InChIKey |
HOWBSMILMYIFKQ-OBCLEYQXSA-N
|
|||||
| Pharmaceutical Properties |
Molecule Weight
|
1253.434
|
Polar area
|
498.15
|
||
|
Complexity
|
1252.646525
|
xlogp Value
|
-3.68013
|
|||
|
Heavy Count
|
90
|
Rot Bonds
|
35
|
|||
|
Hbond acc
|
16
|
Hbond Donor
|
18
|
|||
The Activity Data of This Peptide
| Peptide Activity Information 1 | [1] | |||||
| IC50 | 10.5±0.2 nM | |||||
|---|---|---|---|---|---|---|
| Binding Affinity Assay |
Radioiodination of D-Tyr6-His5-GnRH, preparation of membrane homogenates from HEK 293 cells stably expressing the GnRH-R and binding of the GnRH-gemcitabine conjugates to the human GnRH-R was performed as described before [43,44]. In brief, aliquots of diluted membrane suspension (50 μL) were added into tubes containing buffer B (25 mM HEPES containing 1 mM CaCl2, 10 mM MgCl2, 0.5% BSA, pH 7.4 at 4 °C) and 100,000-120,000 cpm [125I]-D-Tyr6-His5-GnRH with or without increasing concentrations of GnRH-gemcitabine conjugates in a final volume of 0.2 mL. The mixtures were incubated at 4 °C for 16-19 h and then filtered using a Brandel cell harvester through Whatman GF/C glass fiber filters, presoaked for 1-2 h in 0.5% polyethylenimine at 4 °C. The filters were washed four times with 1.5 mL of ice-cold 50 mM Tris-HCl, pH 7.4 at 4 °C. Filters were assessed for radioactivity in a gamma counter (LKB Wallac 1275 minigamma, 80% efficiency). The amount of membrane used was adjusted to ensure that the specific binding was always equal to or less than 10% of the total concentration of the added radioligand. Specific [125I]-D-Tyr6-His5-GnRH binding was defined as total binding less nonspecific binding in the presence of 1000 nM triptorelin (Bachem, Germany). Data for competition binding were analyzed by nonlinear regression analysis, using GraphPad Prism 4.0 (GraphPad Software, San Diego, CA). IC50 values were obtained by fitting the data from competition studies to a one-site competition model.
Click to Show/Hide
|
|||||
| Experimental Condition | HEK 293 cells | |||||
Each Peptide-drug Conjugate Related to This Peptide
Full Information of The Activity Data of The PDC(s) Related to This Peptide
GOXG2 [Investigative]
Identified from the Human Clinical Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
5%
|
|||
| Administration Time | 60 min | ||||
| Administration Dosage | 1 μg/ml | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
GOXG2 was the least stable, followed by GOXG1, while GN4OXG was the most stable.
|
||||
| In Vivo Model | Human plasma. | ||||
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
0%
|
|||
| Administration Time | 10 h | ||||
| Administration Dosage | 1 μM | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
This experiment showed that the more labile pro-drug was GOXG2, which was almost fully degraded in less than 10 hours while GN4OXG and GOXG1 showed similar, enhanced stability compared to GOXG2.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
494 ± 93 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
675 ± 82 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
GOXG1 [Investigative]
Identified from the Human Clinical Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
30%
|
|||
| Administration Time | 60 min | ||||
| Administration Dosage | 1 μg/ml | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
GOXG2 was the least stable, followed by GOXG1, while GN4OXG was the most stable.
|
||||
| In Vivo Model | Human plasma. | ||||
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
10%
|
|||
| Administration Time | 10 h | ||||
| Administration Dosage | 1 μM | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
This experiment showed that the more labile pro-drug was GOXG2, which was almost fully degraded in less than 10 hours while GN4OXG and GOXG1 showed similar, enhanced stability compared to GOXG2.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
611 ± 80 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
754 ± 142 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
GN4OXG [Investigative]
Identified from the Human Clinical Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
75%
|
|||
| Administration Time | 60 min | ||||
| Administration Dosage | 1 μg/ml | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
GOXG2 was the least stable, followed by GOXG1, while GN4OXG was the most stable.
|
||||
| In Vivo Model | Human plasma. | ||||
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Relative intensity |
12%
|
|||
| Administration Time | 10 h | ||||
| Administration Dosage | 1 μM | ||||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
This experiment showed that the more labile pro-drug was GOXG2, which was almost fully degraded in less than 10 hours while GN4OXG and GOXG1 showed similar, enhanced stability compared to GOXG2.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
590 ± 62 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Prostate cancer | ||||
| Efficacy Data | Half maximal inhibitory concentration (IC50) |
833 ± 27 nM
|
|||
| MOA of PDC |
We aimed to construct PDCs with linker controllable drug release rates simply by manipulating the linker unit. For a more rapid drug-release rate we developed GOXG1 and GOXG2. These conjugates bear a carboxylate ester linker directly attached to the primary and the secondary alcohol group of the drug respectively, followed by oxime and amide bond. The primary alcohol of gemcitabine has been used since it is involved in the phosphorylation process, through which gemcitabine exerts its cytotoxic effect. Therefore, we aimed to block the primary alcohol and examine its effect (GOXG1) and also take advantage of the secondary alcohol which could lead to a PDC with a completely different profile (GOXG2), although they share structural similarities. For a slower drug release rate, we designed and developed the PDC GN4OXG that contains an amide bond on the 4-N position of the parent drug followed by click oxime ligation and another amide bond. The stability of this molecule should be enhanced since it is devoid of rapidly hydrolyzable ester bonds. Furthermore, in this PDC since the 4-NH2 moiety of gemcitabine is capped it could further surmount the rapid gemcitabine metabolism that leads to the formation of dFdU, after the enzymatic 4-N deamination of gemcitabine by cytidine deaminase (CDA).
Click to Show/Hide
|
||||
| Description |
The most potent among them was found to be pro-drug GOXG2 with IC50 494 ± 93 nM and 675 ± 82 nM against DU145 and PC3, respectively. Regarding the other two conjugates, GN4OXG was found to be the next most cytotoxic compound against DU145 with IC50 590 ± 62 nM, followed by the least toxic GOXG1 with IC50 611 ± 80 nM. In contrast, GOXG1 was the second more toxic against PC3 with IC50 754 ± 142 nM, followed by the least toxic GN4OXG which showed IC50 833 ± 27 nM. The results are summarized in Table 2.
Click to Show/Hide
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
D-Lys6-GnRH-gemcitabine(2G2) [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
0.11 nM
|
|||
| Description |
The presented data show that 2G2, 2G1 and GSHG bind to GnRH-R with 95.5-, 15.2-, and 4.4-fold higher affinity, respectively, than that of the native peptide D-Lys6-GnRH (10.5 ± 0.2 nM, according to our former study [3]).
|
||||
| In Vitro Model | Normal | HEK293 cell | CVCL_0045 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
449.1 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Invasive breast carcinoma | MCF-7 cell | CVCL_0031 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
9761 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 40000 nM | |||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 40000 nM | |||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
D-Lys6-GnRH-gemcitabine(2G1) [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
0.69 nM
|
|||
| Description |
The presented data show that 2G2, 2G1 and GSHG bind to GnRH-R with 95.5-, 15.2-, and 4.4-fold higher affinity, respectively, than that of the native peptide D-Lys6-GnRH (10.5 ± 0.2 nM, according to our former study [3]).
|
||||
| In Vitro Model | Normal | HEK293 cell | CVCL_0045 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
621.3 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Invasive breast carcinoma | MCF-7 cell | CVCL_0031 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 40000 nM | |||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 40000 nM | |||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 40000 nM | |||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
D-Lys6-GnRH-gemcitabine(GSHG) [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
2.40 nM
|
|||
| Description |
The presented data show that 2G2, 2G1 and GSHG bind to GnRH-R with 95.5-, 15.2-, and 4.4-fold higher affinity, respectively, than that of the native peptide D-Lys6-GnRH (10.5 ± 0.2 nM, according to our former study [3]).
|
||||
| In Vitro Model | Normal | HEK293 cell | CVCL_0045 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
55.5 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Invasive breast carcinoma | MCF-7 cell | CVCL_0031 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
684 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | DU145 cell | CVCL_0105 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
937 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Prostate carcinoma | PC-3 cell | CVCL_0035 | ||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Tumor | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
2387 nM
|
|||
| Description |
GSHGpossesses the highest cytotoxic effect among the three conjugates, which is comparable with that of gemcitabine in the examined cell lines and especially regarding MCF-7 cells.
|
||||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
AEZS-108 [Terminated in Phase 3]
Identified from the Human Clinical Data
| Experiment 1 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Vomit |
32%
|
|||
| Description |
The most common all-grade adverse events were hematologic (22; 88%), fatigue (19; 76%), anorexia (13; 52%),alopecia(13; 52%), nausea (13; 52%), and vomiting (8;32%).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 2 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Neutropenia |
56%
|
|||
| Description |
The most common hematologic adverse event was neutropenia at 56% (all grades).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 3 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Nausea toxicity |
52%
|
|||
| Description |
The most common all-grade adverse events were hematologic (22; 88%), fatigue (19; 76%), anorexia (13; 52%),alopecia(13; 52%), nausea (13; 52%), and vomiting (8;32%).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 4 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Median progression-free survival (mPFS) |
3.8 months
|
|||
| Description |
With a median follow-up of 16.1 months (range, 3.2-36.1), the median PFS was 3.8 months (95% confidence interval [CI], 2.1-4.4) and median OS was 6.0 months (95% CI, 4.2-10.1; Figure 3).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 5 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Median overall survival (mOS) |
6.0 months
|
|||
| Description |
With a median follow-up of 16.1 months (range, 3.2-36.1), the median PFS was 3.8 months (95% confidence interval [CI], 2.1-4.4) and median OS was 6.0 months (95% CI, 4.2-10.1; Figure 3).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 6 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Hematologic toxicity |
88%
|
|||
| Description |
The most common all-grade adverse events were hematologic (22; 88%), fatigue (19; 76%), anorexia (13; 52%),alopecia(13; 52%), nausea (13; 52%), and vomiting (8;32%).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 7 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Grade 3/4 nonhematologic toxicities |
24%
|
|||
| Description |
Six of 25 patients (24%) experienced Grade 3 or 4 nonhematologic toxicities
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 8 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Grade ≥ 3 hematologic toxicity |
56%
|
|||
| Description |
Fourteen of 25 patients (56%) experienced a Grade ≥ 3 hematologic toxicity
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 9 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Fatigue |
76%
|
|||
| Description |
The most common all-grade adverse events were hematologic (22; 88%), fatigue (19; 76%), anorexia (13; 52%),alopecia(13; 52%), nausea (13; 52%), and vomiting (8;32%).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
| Experiment 10 Reporting the Activity Data of This PDC | [3] | ||||
| Indication | Castration and taxane resistant prostate cancer | ||||
| Efficacy Data | Alopecia toxicity |
52.00%
|
|||
| Description |
The most common all-grade adverse events were hematologic (22; 88%), fatigue (19; 76%), anorexia (13; 52%),alopecia(13; 52%), nausea (13; 52%), and vomiting (8;32%).
|
||||
| In Vivo Model | Men with histologically confirmed prostatic adenocarcinoma. | ||||
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
0%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 5 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 2 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
5%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 1 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 3 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
6%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 2.5 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 4 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
25%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 5 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM3DOX320 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 5 Reporting the Activity Data of This PDC | [5] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
36.30%
|
|||
| Administration Time | 24 h | ||||
| Administration Dosage | 5 µM | ||||
| Evaluation Method | MTS assay | ||||
| MOA of PDC |
Our results show that AEZS-108 upregulates the expression of MASPIN/SERPINB5 tumor suppressor gene, which is downregulated in normal uvea and UM specimens independently from the LHRH receptor-ligand interaction. AEZS-108 also substantially downregulates hypoxia-inducible factor 1 alpha (HIF1A) expression.
|
||||
| Description |
In order to investigate whether AEZS-108 inhibits cell proliferation and its extent, OCM3 cells were treated either with 5 M AEZS-108 or equal amount of doxorubicin. MTS assay was performed after 24 and 48 hours of treatment. AEZS-108 and doxorubicin have been shown to reduce cell proliferation by 36.3% (p< 0.001) and 62.9% (p< 0.001) respectively after 24 hours, and by 84.7% (p< 0.001) and 89.7% (p< 0.001) respectively after 48 hours, (Figure 2).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Experiment 6 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
48%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 2.5 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM3DOX320 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 7 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
80%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 1 µM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM3DOX320 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 8 Reporting the Activity Data of This PDC | [5] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
84.70%
|
|||
| Administration Time | 48 h | ||||
| Administration Dosage | 5 µM | ||||
| Evaluation Method | MTS assay | ||||
| MOA of PDC |
Our results show that AEZS-108 upregulates the expression of MASPIN/SERPINB5 tumor suppressor gene, which is downregulated in normal uvea and UM specimens independently from the LHRH receptor-ligand interaction. AEZS-108 also substantially downregulates hypoxia-inducible factor 1 alpha (HIF1A) expression.
|
||||
| Description |
In order to investigate whether AEZS-108 inhibits cell proliferation and its extent, OCM3 cells were treated either with 5 M AEZS-108 or equal amount of doxorubicin. MTS assay was performed after 24 and 48 hours of treatment. AEZS-108 and doxorubicin have been shown to reduce cell proliferation by 36.3% (p< 0.001) and 62.9% (p< 0.001) respectively after 24 hours, and by 84.7% (p< 0.001) and 89.7% (p< 0.001) respectively after 48 hours, (Figure 2).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Experiment 9 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
105%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 320 nM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM3DOX320 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 10 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
120%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 40 nM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM3DOX320 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
| Experiment 11 Reporting the Activity Data of This PDC | [4] | ||||
| Indication | Uveal melanoma | ||||
| Efficacy Data | Cell viability |
130%
|
|||
| Administration Time | 72 h | ||||
| Administration Dosage | 40 nM | ||||
| Evaluation Method | MTT assay | ||||
| Description |
OCM3DOX320cells did not show significant difference in cell viability in the presence of 1 uM DOX when compared to untreated control cells. 1 uM DOX induced significant cell death of OCM3 cells but did not cause significant cell death in OCM3DOX320cells, confirming DOX resistance. AN-152 at lower concentration (40 nM, 320 nM) increased cell proliferation significantly compared to equimolar dose of DOX which does not have any effect on cell viability at this concentration in OCM3 cells. However, higher concentrations of AN-152 can effectively inhibit cell proliferation in both cell lines. Higher concentrations (1-5 uM) of DOX and AN-152 showed no significantly different effect either on OCM3 or OCM3DOX320cells (Fig. 3).
Click to Show/Hide
|
||||
| In Vitro Model | Cutaneous melanoma | OCM-3 cell | CVCL_6937 | ||
| Half life period | 2 h | ||||
References