Peptide Information
General Information of This Peptide
| Peptide ID |
PEP00158
|
|||||
|---|---|---|---|---|---|---|
| Peptide Name |
[K4,F7,K22,P34]-NPY
|
|||||
| Structure |
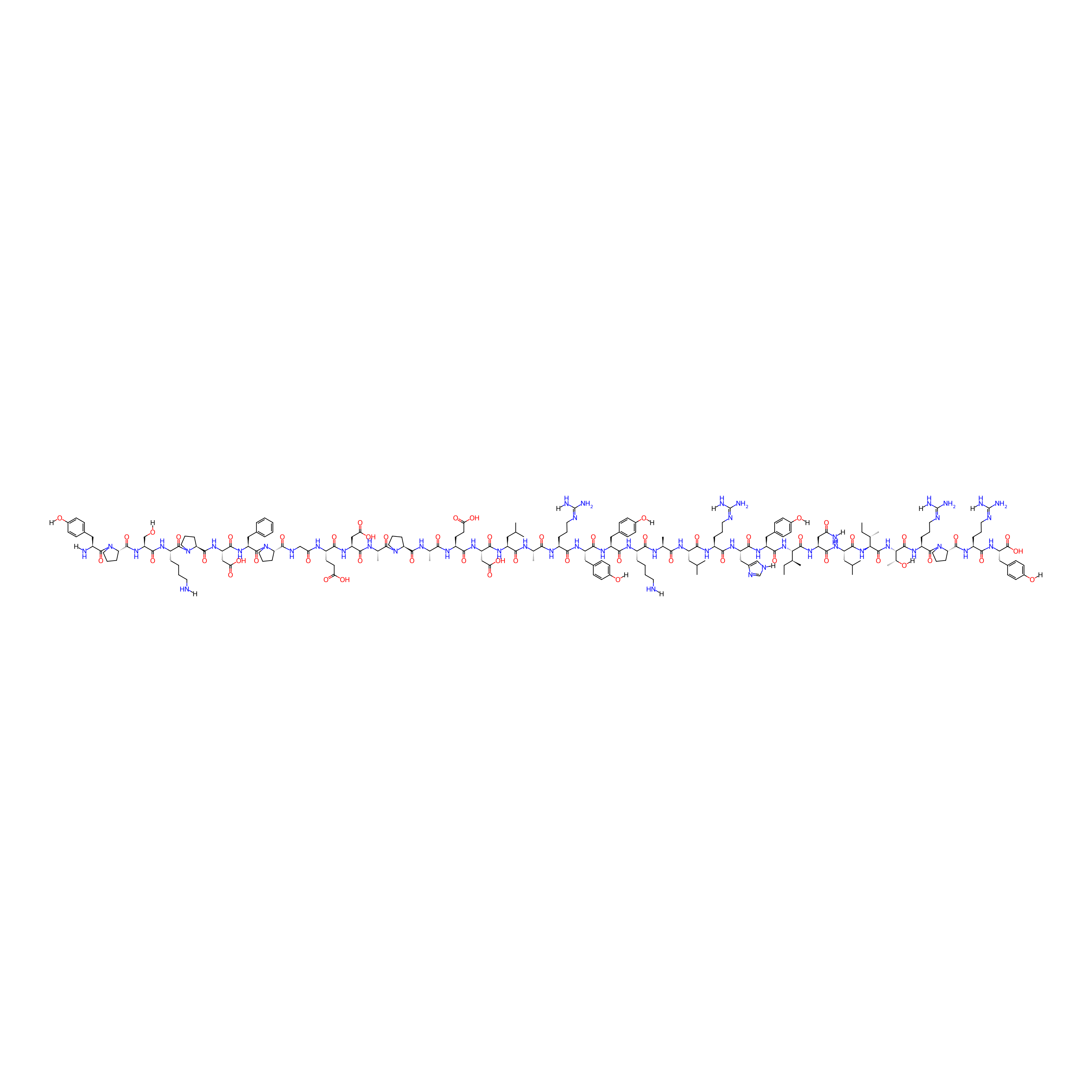
|
|||||
| Sequence |
YPSKPDFPGEDAPAEDLARYYKALRHYINLITRPRY
|
|||||
| Peptide Type |
Linear
|
|||||
| Receptor Name |
Neuropeptide Y receptor type 1 (NPY1R)
|
Receptor Info | ||||
| PDC Transmembrane Types | Cell targeting peptides (CTPs) | |||||
| Formula |
C198H295N53O55
|
|||||
| Isosmiles |
[H]NCCCC[C@H](NC(=O)[C@H](Cc1ccc(O[H])cc1)NC(=O)[C@H](Cc1ccc(O[H])cc1)NC(=O)[C@H](CCC/N=C(/N)N[H])NC(=O)[C@H](C)NC(=O)[C@H](CC(C)C)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](C)NC(=O)[C@@H]1CCCN1C(=O)[C@H](C)NC(=O)[C@H](CC(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)CNC(=O)[C@@H]1CCCN1C(=O)[C@H](Cc1ccccc1)NC(=O)[C@H](CC(=O)O)NC(=O)[C@@H]1CCCN1C(=O)[C@H](CCCCN[H])NC(=O)[C@H](CO[H])NC(=O)[C@@H]1CCCN1C(=O)[C@H](Cc1ccc(O[H])cc1)N[H])C(=O)N[C@@H](C)C(=O)N[C@@H](CC(C)C)C(=O)N[C@@H](CCC/N=C(/N)N[H])C(=O)N[C@@H](Cc1cn([H])cn1)C(=O)N[C@@H](Cc1ccc(O[H])cc1)C(=O)N[C@]([H])(C(=O)N[C@@H](CC(=O)N[H])C(=O)N[C@@H](CC(C)C)C(=O)N[C@]([H])(C(=O)N[C@]([H])(C(=O)N[C@@H](CCC/N=C(/N)N[H])C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC/N=C(/N)N[H])C(=O)N[C@@H](Cc1ccc(O[H])cc1)C(=O)O)[C@@H](C)O[H])[C@@H](C)CC)[C@@H](C)CC
|
|||||
| InChI |
InChI=1S/C198H295N53O55/c1-16-103(9)157(186(297)240-138(92-150(202)259)175(286)232-133(84-102(7)8)178(289)245-158(104(10)17-2)187(298)246-159(109(15)253)188(299)229-130(41-28-76-215-198(209)210)192(303)250-80-32-45-148(250)183(294)227-126(40-27-75-214-197(207)208)166(277)242-143(194(305)306)90-115-55-65-121(258)66-56-115)244-179(290)136(88-114-53-63-120(257)64-54-114)235-174(285)137(91-116-96-211-99-217-116)236-165(276)125(39-26-74-213-196(205)206)226-171(282)132(83-101(5)6)230-162(273)106(12)218-163(274)123(36-21-23-71-199)225-172(283)134(86-112-49-59-118(255)60-50-112)234-173(284)135(87-113-51-61-119(256)62-52-113)233-164(275)124(38-25-73-212-195(203)204)223-160(271)105(11)219-169(280)131(82-100(3)4)231-176(287)140(94-155(267)268)238-168(279)128(68-70-153(263)264)224-161(272)107(13)220-182(293)146-43-30-77-247(146)189(300)108(14)221-170(281)139(93-154(265)266)237-167(278)127(67-69-152(261)262)222-151(260)97-216-181(292)145-42-29-79-249(145)193(304)142(89-110-34-19-18-20-35-110)241-177(288)141(95-156(269)270)239-184(295)149-46-33-81-251(149)191(302)129(37-22-24-72-200)228-180(291)144(98-252)243-185(296)147-44-31-78-248(147)190(301)122(201)85-111-47-57-117(254)58-48-111/h18-20,34-35,47-66,96,99-109,122-149,157-159,252-258H,16-17,21-33,36-46,67-95,97-98,199-201H2,1-15H3,(H2,202,259)(H,211,217)(H,216,292)(H,218,274)(H,219,280)(H,220,293)(H,221,281)(H,222,260)(H,223,271)(H,224,272)(H,225,283)(H,226,282)(H,227,294)(H,228,291)(H,229,299)(H,230,273)(H,231,287)(H,232,286)(H,233,275)(H,234,284)(H,235,285)(H,236,276)(H,237,278)(H,238,279)(H,239,295)(H,240,297)(H,241,288)(H,242,277)(H,243,296)(H,244,290)(H,245,289)(H,246,298)(H,261,262)(H,263,264)(H,265,266)(H,267,268)(H,269,270)(H,305,306)(H4,203,204,212)(H4,205,206,213)(H4,207,208,214)(H4,209,210,215)/t103-,104-,105-,106-,107-,108-,109+,122-,123-,124-,125-,126-,127-,128-,129-,130-,131-,132-,133-,134-,135-,136-,137-,138-,139-,140-,141-,142-,143-,144-,145-,146-,147-,148-,149-,157-,158-,159-/m0/s1
|
|||||
| InChIKey |
VKXHTGBMWJGPHJ-VUZUKZSBSA-N
|
|||||
| Pharmaceutical Properties |
Molecule Weight
|
4297.854
|
Polar area
|
1747.39
|
||
|
Complexity
|
4295.191611
|
xlogp Value
|
-13.3366
|
|||
|
Heavy Count
|
306
|
Rot Bonds
|
140
|
|||
|
Hbond acc
|
57
|
Hbond Donor
|
56
|
|||
The Activity Data of This Peptide
| Peptide Activity Information 1 | [1] | |||||
| EC50 | 5 nM | |||||
|---|---|---|---|---|---|---|
| Binding Affinity Assay |
Signal transduction inositol phosphate (IP) accumulation assay was performed like described before.(40)Briefly, stably transfected COS-7 cells were seeded into 48-well plates (50000 cells per well) and cultured overnight. After labeling cells with 2 μCi/mL myo-[2-3H]-inositol in culture medium without penicillin and streptomycin, cells were washed and incubated with peptide concentrations ranging from 10-5to 10-12M in DMEM supplied with 10 mM LiCl for 1 h at 37 °C. Radioactive IP species were isolated by anion exchange chromatography and measured in a scintillation counter. EC50and pEC50values were calculated from concentration-response curves by using nonlinear regression (GraphPad Prism 5.0). Assays were performed in duplicate in at least two independent experiments.
Click to Show/Hide
|
|||||
| Experimental Condition | COS-7 cells; hY<sub>1</sup>R | |||||
| Peptide Activity Information 2 | [1] | |||||
| EC50 | 7.1±0.1 nM | |||||
| Binding Affinity Assay |
Signal transduction inositol phosphate (IP) accumulation assay was performed like described before.(40)Briefly, stably transfected COS-7 cells were seeded into 48-well plates (50000 cells per well) and cultured overnight. After labeling cells with 2 μCi/mL myo-[2-3H]-inositol in culture medium without penicillin and streptomycin, cells were washed and incubated with peptide concentrations ranging from 10-5to 10-12M in DMEM supplied with 10 mM LiCl for 1 h at 37 °C. Radioactive IP species were isolated by anion exchange chromatography and measured in a scintillation counter. EC50and pEC50values were calculated from concentration-response curves by using nonlinear regression (GraphPad Prism 5.0). Assays were performed in duplicate in at least two independent experiments.
Click to Show/Hide
|
|||||
| Experimental Condition | COS-7 cells; hY<sub>2</sup>R | |||||
| Peptide Activity Information 3 | [1] | |||||
| EC50 | 8.3±0.1 nM | |||||
| Binding Affinity Assay |
Signal transduction inositol phosphate (IP) accumulation assay was performed like described before.(40)Briefly, stably transfected COS-7 cells were seeded into 48-well plates (50000 cells per well) and cultured overnight. After labeling cells with 2 μCi/mL myo-[2-3H]-inositol in culture medium without penicillin and streptomycin, cells were washed and incubated with peptide concentrations ranging from 10-5to 10-12M in DMEM supplied with 10 mM LiCl for 1 h at 37 °C. Radioactive IP species were isolated by anion exchange chromatography and measured in a scintillation counter. EC50and pEC50values were calculated from concentration-response curves by using nonlinear regression (GraphPad Prism 5.0). Assays were performed in duplicate in at least two independent experiments.
Click to Show/Hide
|
|||||
| Experimental Condition | COS-7 cells; hY<sub>1</sup>R | |||||
| Peptide Activity Information 4 | [1] | |||||
| EC50 | 87 nM | |||||
| Binding Affinity Assay |
Signal transduction inositol phosphate (IP) accumulation assay was performed like described before.(40)Briefly, stably transfected COS-7 cells were seeded into 48-well plates (50000 cells per well) and cultured overnight. After labeling cells with 2 μCi/mL myo-[2-3H]-inositol in culture medium without penicillin and streptomycin, cells were washed and incubated with peptide concentrations ranging from 10-5to 10-12M in DMEM supplied with 10 mM LiCl for 1 h at 37 °C. Radioactive IP species were isolated by anion exchange chromatography and measured in a scintillation counter. EC50and pEC50values were calculated from concentration-response curves by using nonlinear regression (GraphPad Prism 5.0). Assays were performed in duplicate in at least two independent experiments.
Click to Show/Hide
|
|||||
| Experimental Condition | COS-7 cells; hY<sub>2</sup>R | |||||
Each Peptide-drug Conjugate Related to This Peptide
Full Information of The Activity Data of The PDC(s) Related to This Peptide
Cytolysin[F7,P34]-NPY bioconjugate [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
236.1 nM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-468 cell | CVCL_0419 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [2] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
405.3 nM
|
|||
| In Vitro Model | Invasive breast carcinoma | MCF-7 cell | CVCL_0031 | ||
PDC-5d [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
2.0 (5.7 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-468 cell | CVCL_0419 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
4.2 (5.4 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
5.0 (5.3 ± 0.1) μM
|
|||
| In Vitro Model | Invasive breast carcinoma | T-47D cell | CVCL_0553 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
8.9 nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
PDC-5b [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
3.6 (5.4 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-468 cell | CVCL_0419 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
7.7 (5.1 ± 0.1) μM
|
|||
| In Vitro Model | Invasive breast carcinoma | T-47D cell | CVCL_0553 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
8.1 (5.1 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
3.0 (8.5 ± 0.1) nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
226 (6.6 ± 0.1) nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
PDC-5c [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
3.8 (5.4 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
4.0 (5.4 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-468 cell | CVCL_0419 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 10 µM | |||
| In Vitro Model | Invasive breast carcinoma | T-47D cell | CVCL_0553 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
2.8 (8.5 ± 0.1) nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
PDC-5a [Investigative]
Revealed Based on the Cell Line Data
| Experiment 1 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) |
18 (4.8 ± 0.1) μM
|
|||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-468 cell | CVCL_0419 | ||
| Experiment 2 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 25 µM | |||
| In Vitro Model | Invasive breast carcinoma | T-47D cell | CVCL_0553 | ||
| Experiment 3 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Inhibitory Concentration (IC50) | > 25 µM | |||
| In Vitro Model | Breast adenocarcinoma | MDA-MB-231 cell | CVCL_0062 | ||
| Experiment 4 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
2.9 (8.5 ± 0.1) nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
| Experiment 5 Reporting the Activity Data of This PDC | [1] | ||||
| Indication | Breast cancer | ||||
| Efficacy Data | Half Maximal Effective Concentration (EC50) |
209 (6.7 ± 0.1) nM
|
|||
| In Vitro Model | Normal | COS-7 cell | CVCL_0224 | ||
References